After UvrA has dissociated, UvrB (at the damaged site) forms a complex with UvrC. This is an autocatalytic reaction, since it is catalyzed by UvrA, which is itself an ATPase. UvrA 2 then dissociates, in a step that requires ATP hydrolysis. UvrA 2 (a dimer) and Uvr B recognize the damaged site as a (UvrA)2UvrB complex. In more detail, the process goes as follows (Figure 7.14). Since the undamaged template directs the synthesis by DNA polymerase, the resulting duplex DNA is no longer damaged. This can be filled in by DNA polymerase and the remaining nick sealed by DNA ligase. 
After the damaged segment has been excised, a gap of 12 to 13 nucleotides remains in the DNA. Thus the UvrABC dynamic complex and the UvrBC complex can be called excinucleases. The two cuts around the damage allow the single-stranded segment containing the damage to be excised by the helicase activity of UvrD. The UvrABC proteins form a dynamic complex that recognizes damage and makes endonucleolytic cuts on both sides. This leaves a gapped substrate for copying by DNA polymerase and pasting by DNA ligase. Thus for this system, the UvrABC and UvrD proteins carry out a series of steps in the cutting phase of excision repair. Then UvrD (also called helicase II), the product of the uvrDgene, unwinds the DNA, releasing the damaged segment. It then cleaves on both sides of the damage. 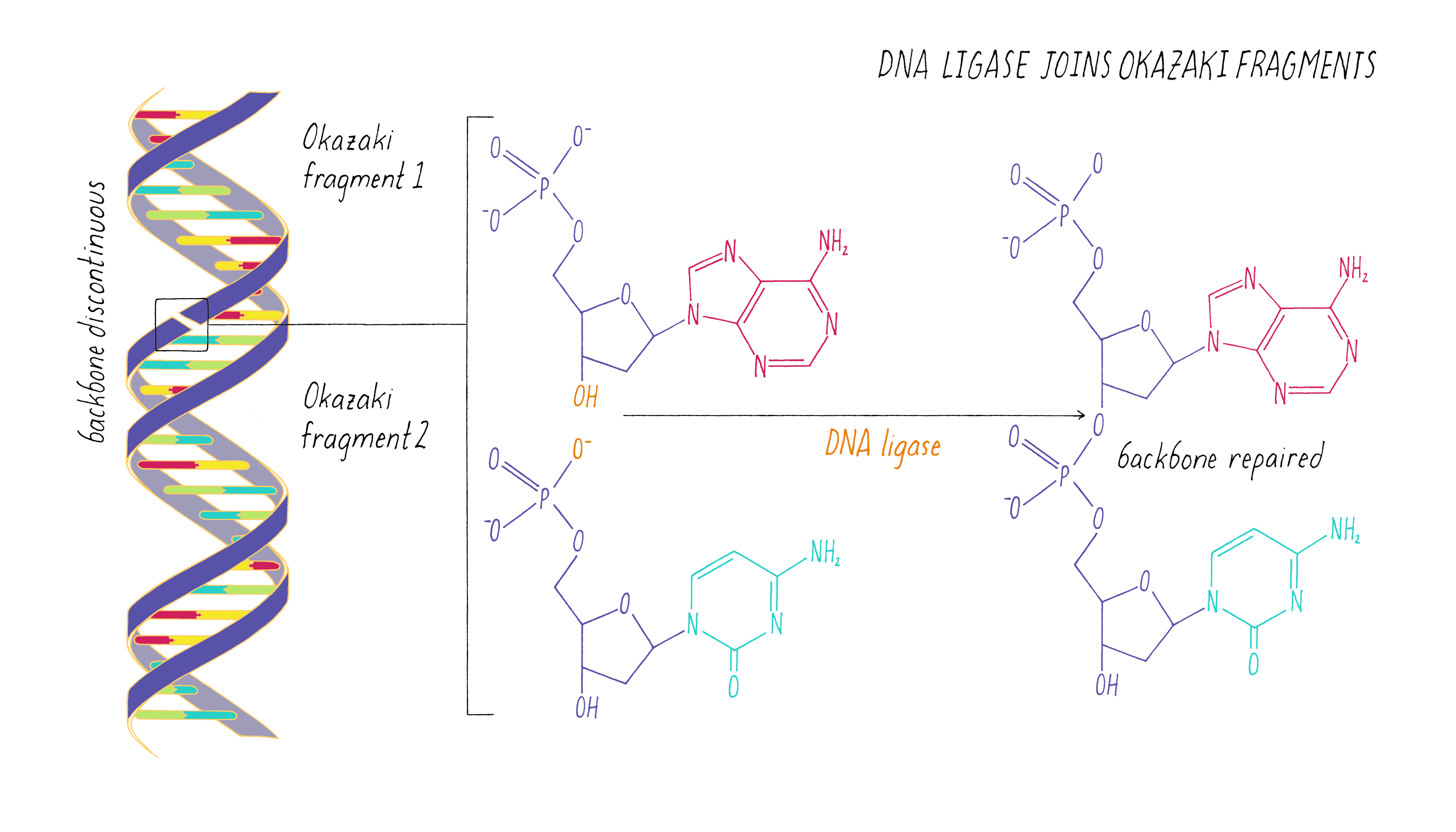
The UvrABC complex recognizes damage-induced structural distortions in the DNA, such as pyrimidine dimers. UvrA is the protein encoded by uvrA, UvrB is encoded by uvrB, and so on. The polypeptide products of the uvrA, uvrB, and uvrCgenes are subunits of a multisubunit enzyme called the UvrABC excinuclease. The enzymes encoded by the uvrgenes have been studied in detail. Mutant strains that are more sensitive to UV damage are defective in the genes that confer UV- resistance, i.e. Samples of each irradiated culture are then plated and the number of surviving colonies are counted (plotted as a logarithmic function on the vertical axis). Cultures of bacteria are exposed to increasing doses of UV radiation, plotted along the horizontal axis. Survival curve of bacteria exposed to UV radiation. Four of the complementation groups, or genes, encode proteins that play major rules in NER they are uvrA, uvrB, uvrCand uvrD. By collecting large numbers of such mutants and testing them for their ability to restore resistance to UV radiation in combination, complementation groups were identified. Mutant strains can be identified that are substantially more sensitive to UV radiation these are defective in the functions needed for UV- resistance, abbreviated uvr. coli cells are killed only at higher doses of UV radiation. As illustrated in Figure 7.13, wild type E. The genes encoding this repair function were discovered as mutants that are highly sensitive to UV damage, indicating that the mutants are defective in UV repair. Some of the best-characterized enzymes catalyzing this process are the UvrABC excinuclease and the UvrD helicase in E. NER occurs in almost all organisms examined. The common feature of damage that is repaired by nucleotide excision is that the modified nucleotides cause a significant distortion in the DNA helix. This repair system is used to remove pyrimidine dimers formed by UV radiation as well as nucleotides modified by bulky chemical adducts. In nucleotide excision repair, damaged bases are cut out within a string of nucleotides, and replaced with DNA as directed by the undamaged template strand. The methylated enzyme is no longer active, hence this has been referred to as a suicide mechanism for the enzyme. It then removes the methyl group, transferring it to an amino acid of the enzyme. coli, recognizes O6‑methylguanine in duplex DNA. For instance, the enzyme O6‑methylguanine methyltransferase, encoded by the adagene in E. coli.Ī second example of the reversal of damage is the removal of methyl groups. The photolyase enzyme has two subunits, which are encoded by the phrA and phrBgenes in E. However, the result is that the DNA structure has been returned to its state prior to damage by UV. Note that this is not formally the reverse of the reaction that formed the pyrimidine dimers, since energy from visible light is used to break the bonds between the pyrimidines, and no UV radiation is released. The enzyme photolyase binds to a pyrimidine dimer and catalyzes a second photochemical reaction (this time using visible light) that breaks the cyclobutane ring and reforms the two adjacent thymidylates in DNA. Photoreactivation is a light-dependent process used by bacteria to reverse pyrimidine dimers formed by UV radiation. This occurs by specific enzyme systems recognizing the altered base and breaking bonds to remove the adduct or change the base back to its normal structure. Some kinds of covalent alteration to bases in DNA can be directly reversed.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
